piggybac vector ppbcag ntom20 moxbfp gs lov2 ires puro (Addgene inc)
Structured Review
Piggybac Vector Ppbcag Ntom20 Moxbfp Gs Lov2 Ires Puro, supplied by Addgene inc, used in various techniques. Bioz Stars score: 93/100, based on 4 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more
https://www.bioz.com/result/piggybac vector ppbcag ntom20 moxbfp gs lov2 ires puro/product/Addgene inc
Average 93 stars, based on 4 article reviews
Images
1) Product Images from "Optogenetic stimulation of Lbc GEF-mediated Rho activity dynamics promotes cell invasion"
Article Title: Optogenetic stimulation of Lbc GEF-mediated Rho activity dynamics promotes cell invasion
Journal: bioRxiv
doi: 10.1101/2025.03.28.646036

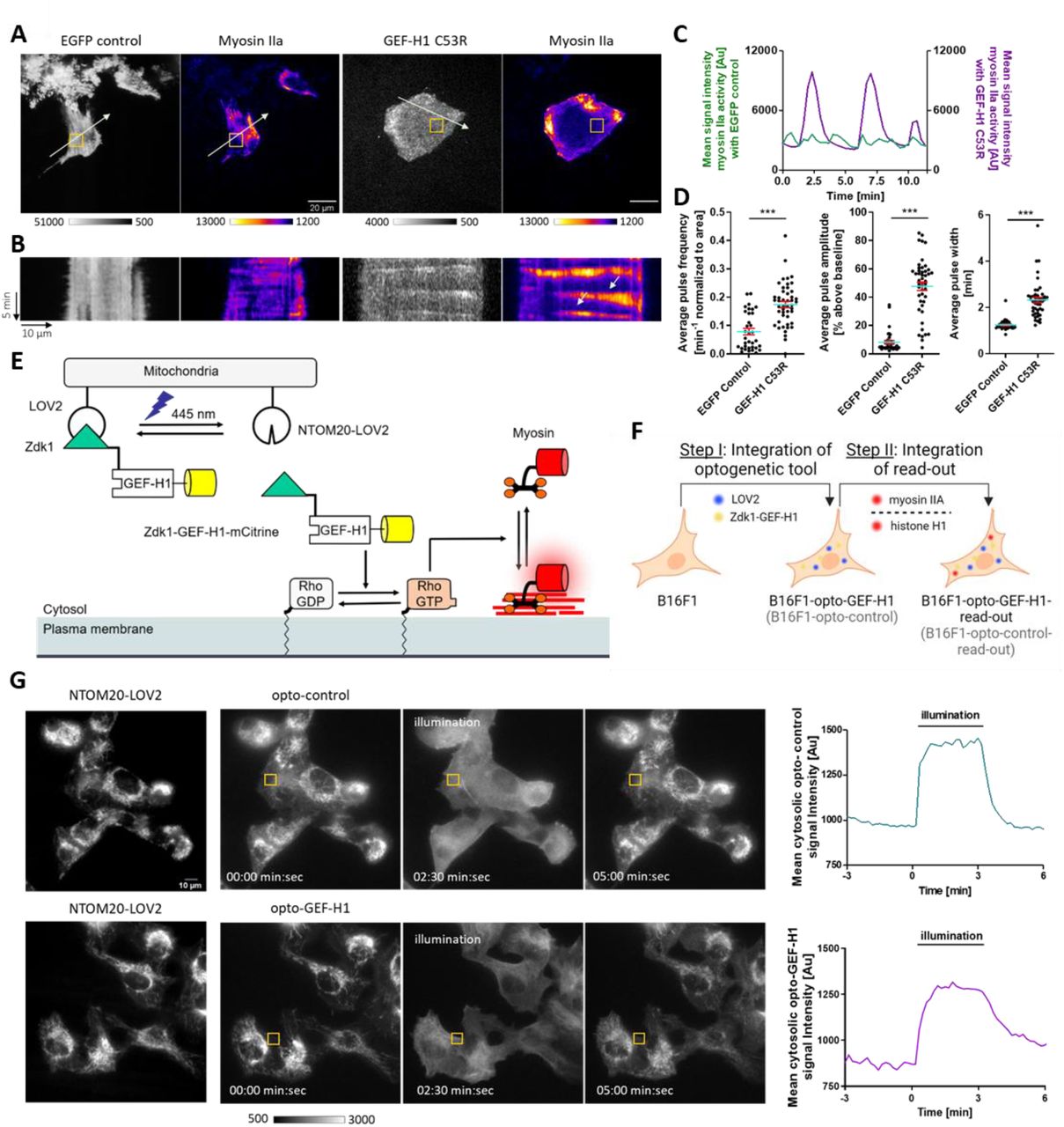
Figure Legend Snippet: (A-D) Increased expression of a constitutively active GEF-H1 mutant in B16F1 mouse melanoma cells leads to enhanced cell contraction dynamics. (A) Representative TIRF images of cells transiently expressing myosin IIa (pCMV-mCherry-MHC IIA) together with the constitutively active GEF-H1 C53R mutant (pCMV5-EGFP-GEF-HI C53R) or a control vector (pEGFP-N1), respectively. Scale bar = 20 μm. (B) Corresponding kymograph analysis along the arrows in A. (C) Myosin IIa signal intensity plots corresponding to the orange box in A. (D) Quantification of average contraction pulse frequency, amplitude and width. N=41 GEF-H1 C53R cells and N=24 control cells from 3 independent experiments, Error bars represent S.E.M., Unpaired t-test. (E) Schematic representation of optogenetic GEF-H1 release from the mitochondria into the cytosol and subsequent stimulation of cell contraction dynamics by myosin IIa. (F) Schematic representation of stepwise strategy to generate stable B16F1 cell lines expressing a GEF-H1-coupled optogenetic tool, the corresponding control and fluorescently tagged read-out proteins. (G) Left: Representative epifluorescence images showing the mitochondrial anchored photo-sensitive LOV2 domain (NTOM20-moxBFP-LOV2) and opto-control (mCitrine-Zdk1), or opto-GEF-H1 (mCitrine-Zdk1-GEF-H1 C53R) before, during and after optogenetic stimulation at 445-488 nm. Scale bar = 10 μm. Right: Intensity plots of opto-control and opto-GEF-H1 signal corresponding to the orange boxes in left panels.
Techniques Used: Expressing, Mutagenesis, Control, Plasmid Preparation




